Jian Hu, Kyle Coleman, Edward B. Lee, Humam Kadara, Linghua Wang, Mingyao Li
Under revision
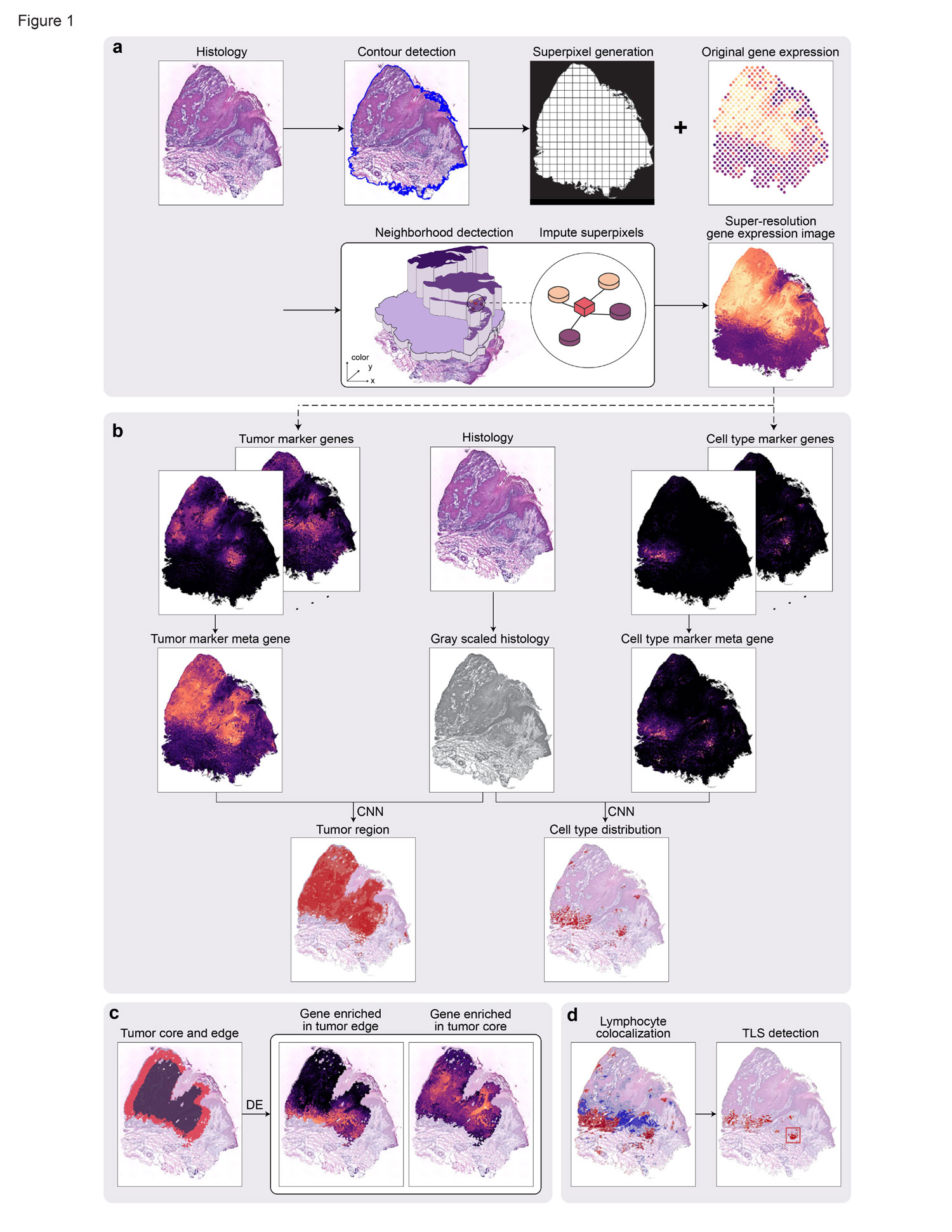
Spatial transcriptomics have enabled the comprehensive characterization of gene expression in tumor microenvironment. However, ST only measures expression in discrete spots, which limits their usefulness in studying the detailed structure of TME. Here we present TESLA, a machine learning framework for multi-level tissue annotation in ST. TESLA integrates histological information with gene expression to annotate heterogeneous immune and tumor cells directly on the histology image, and further detects tertiary lymphoid structures and differential transcriptome programs between the edge and core of a tumor. TESLA provides a powerful tool for understanding the spatial architecture of the TME.
